The Charles Bronfman Institute for Personalized Medicine has been enrolling participants in the BioMe biorepository since 2006. A summary of participants by major ancestral category, and of specimens and data available is shown below.
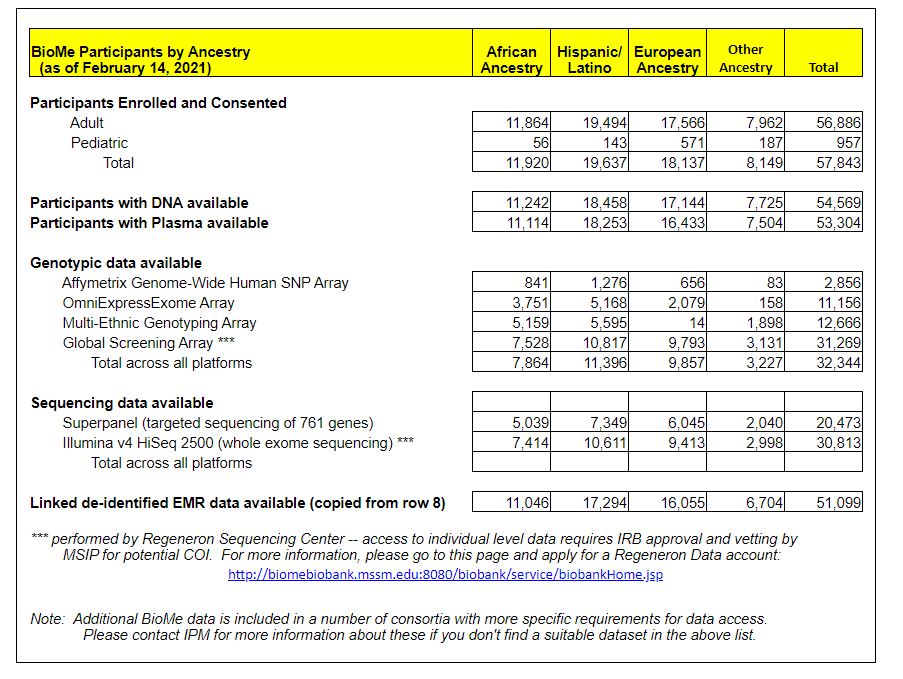
In addition to ancestral diversity, the BioMe biorepository encompasses the full spectrum of traits or diseases seen across the Mount Sinai Health System. A table showing counts of participants with commonly studied traits or diseases is available here (opens in new window):
To request access to any BioMe dataset(s), please go to http://biomebiobank.mssm.edu:8080/biobank/service/ipmHome.jsp
For Regeneron datasets, select Regeneron Data (create an account if you do not have one already) and complete a Data Request. For data from other sources, use the simpler Data Request found under the heading Other than Regeneron Data.